

O estudo da agregação de proteínas abrange uma ampla gama de interações e mecanismos. Muitos estudos nessa área investigam a agregação de proteínas mal enoveladas, que se acredita ser responsável por muitas doenças degenerativas. Como a agregação geralmente leva a uma alteração física no tamanho da proteína, a análise do tamanho de partículas tem se mostrado uma técnica experimental útil nesse campo. Embora a dispersão dinâmica de luz (DLS) seja amplamente utilizada para caracterizar proteínas, em muitos casos os agregados são grandes demais para serem medidos com precisão por essa técnica. O analisador de tamanho de partículas por difração a laser LA-960 possui a capacidade única de medir de 10 nanômetros a 5 milímetros, detectando com precisão os agregados e, em alguns casos, o tamanho da proteína primária.
Para cientistas da área de proteínas, é de particular interesse quantificar o tamanho, o volume e o número de agregados em um determinado material. O analisador de tamanho de partículas LA-960V2 fornece medições precisas do tamanho e do volume de agregados entre 10 nanômetros e 5 milímetros em concentrações ultrabaixas para dispositivos tradicionais de difração a laser.
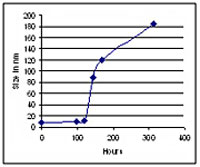
Um exemplo de resultado é mostrado na figura abaixo, utilizando material da FDA (Administração de Alimentos e Medicamentos dos EUA).
Dados de exemplo mostrando o tamanho da proteína primária e agregados acima de 1 mícron. Resultados obtidos com o LA-960.
Diversos tipos de estudos são possíveis com um instrumento de espalhamento dinâmico de luz, como o Analisador de Nanopartículas SZ-100V2. Novamente, como a agregação geralmente pode ser detectada pela alteração do tamanho das partículas, o SZ-100 é útil para agregados menores que um determinado tamanho crítico, no qual o movimento browniano ainda predomina sobre a sedimentação gravitacional. Estudos adicionais podem ser realizados com o SZ-100 para caracterizar como fatores como temperatura, pH, salinidade, surfactante e tempo afetam o enovelamento de proteínas.
A dispersão dinâmica de luz (DLS) é agora amplamente utilizada para estudar a agregação de proteínas, conforme descrito nas notas de aplicação abaixo.
Analisador de nanopartículas
Analisador de Distribuição de Tamanho de Partículas por Dispersão de Laser
Análise simultânea de rastreamento de nanopartículas multiespectrais (NTA)
Você tem alguma dúvida ou solicitação? Utilize este formulário para entrar em contato com nossos especialistas.